Daniel Baer
A Bayesian Linear Mixed Model for Bi-Level Feature Selection
Presenter

Daniel Baer is a postdoctoral fellow in Dr. Sharon Xie's lab. Daniel Baer's research interests include developing models for longitudinal data analysis, feature selection, and measurement error as motivated by complexities arising in the study of neurodegenerative diseases.
Abstract
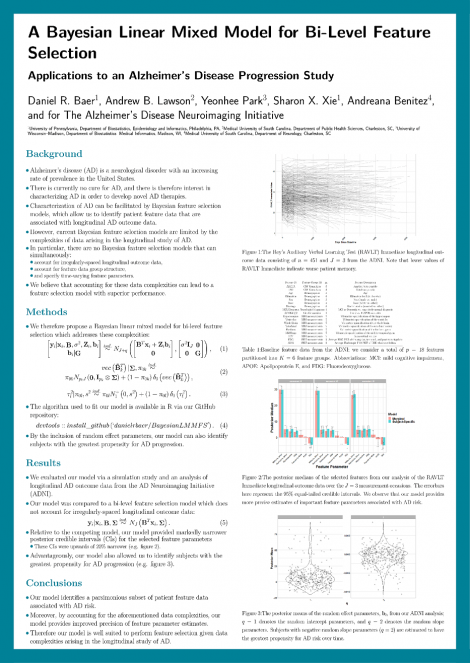
Alzheimer’s disease (AD) is a neurodegenerative disease with an increasing rate of prevalence in the United States. There is currently no cure for AD, and there is therefore interest in characterizing AD in order to develop disease modifying therapies. Characterization of AD can be facilitated by Bayesian feature selection models, which allow us to identify (possibly high-dimensional) patient feature data that are associated with longitudinal AD outcome data (e.g., cognitive scores over time). However, current Bayesian feature selection models are limited by salient complexities arising in the study of longitudinal AD outcome data. In particular, there are no Bayesian feature selection models that can simultaneously account for irregularly-spaced longitudinal outcome data, account for feature data group structure, and specify time-varying feature parameters. Accounting for these complexities can lead to a feature selection model with superior performance. We therefore developed a Bayesian linear mixed model for feature selection which addresses these complexities. We applied our novel approach to analyze longitudinal cognitive scores in the Alzheimer's Disease Neuroimaging Initiative participants with multimodal feature data, including neuroimaging, cerebrospinal fluid biomarkers, genetic markers, neurological diagnoses, and demographics. We found that our model identifies a parsimonious subset of patient feature data associated with rate of cognitive decline, and moreover by accounting for these aforementioned complexities, provides improved precision of feature parameter estimates. Our model therefore represents an effective tool that researchers can use to perform feature selection given complexities arising in the longitudinal study of AD.
Keywords
Feature selection, longitudinal data analysis, Bayesian model, Alzheimer's Disease.About Us
To understand health and disease today, we need new thinking and novel science —the kind we create when multiple disciplines work together from the ground up. That is why this department has put forward a bold vision in population-health science: a single academic home for biostatistics, epidemiology and informatics.
© 2023 Trustees of the University of Pennsylvania. All rights reserved.. | Disclaimer


Comments
Questions: In your comparison model that does not account for irregularly spaced outcomes, does it force a fixed time or is there loss of data? How do the features your model identified compare to the literature? Were there new features?
Thank you for your thoughtful questions.
The competing model discretizes the continuous measurement times associated with the longitudinal outcome data. So there is definitely a loss of data. This is salient as the correlation of longitudinal outcome data is a function of time separation. Therefore a feature selection model which can account for irregularly spaced longitudinal outcome data is advantageous.
The selected features from our analysis of the ADNi data were consistent with the AD literature. For instance, hippocampus volume and CSF TAU were selected as features that were most associated with longitudinal measures of AD risk.
No new features were selected re: modeling the ADNI data. However we found our model was advantageous in terms of providing improved precision re: the selected feature parameter estimates.