Jaesik Kim
HiG2Vec: Hierarchical Representations of Gene Ontology and Genes in the Poincaré Ball
Presenter

I’m a Bioinformatician in Doykoon Kim's lab at the University of Pennsylvania. I obtained a M.S. majoring in Computer Engineering at Ajou University, advised by Prof. Kyung-Ah Sohn. My interests of research are Biomedical Informatics, Machine Learning, Deep Learning, Graph Neural Network, Alzheimer’s Disease, Genome-wide Association Study (GWAS), and Multi-omics Integration.
Abstract
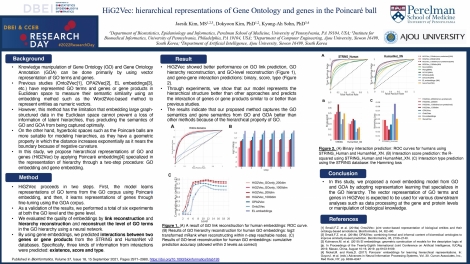
Knowledge manipulation of Gene Ontology (GO) and Gene Ontology Annotation (GOA) can be done primarily by using vector representation of GO terms and genes. Previous studies have represented GO terms and genes or gene products in Euclidean space to measure their semantic similarity using an embedding method such as the Word2Vec-based method to represent entities as numeric vectors. However, this method has the limitation that embedding large graph-structured data in the Euclidean space cannot prevent a loss of information of latent hierarchies, thus precluding the semantics of GO and GOA from being captured optimally. On the other hand, hyperbolic spaces such as the Poincaré balls are more suitable for modeling hierarchies, as they have a geometric property in which the distance increases exponentially as it nears the boundary because of negative curvature. In this study, we propose hierarchical representations of GO and genes (HiG2Vec) by applying Poincaré embedding specialized in the representation of hierarchy through a two-step procedure: GO embedding and gene embedding. Through experiments, we show that our model represents the hierarchical structure better than other approaches and predicts the interaction of genes or gene products similar to or better than previous studies. The results indicate that HiG2Vec is superior to other methods in capturing the GO and gene semantics and in data utilization as well. It can be robustly applied to manipulate various biological knowledge.
Keywords
Machine learning, Representation learning, Gene Ontology, Gene embeddingAbout Us
To understand health and disease today, we need new thinking and novel science —the kind we create when multiple disciplines work together from the ground up. That is why this department has put forward a bold vision in population-health science: a single academic home for biostatistics, epidemiology and informatics.
© 2023 Trustees of the University of Pennsylvania. All rights reserved.. | Disclaimer

